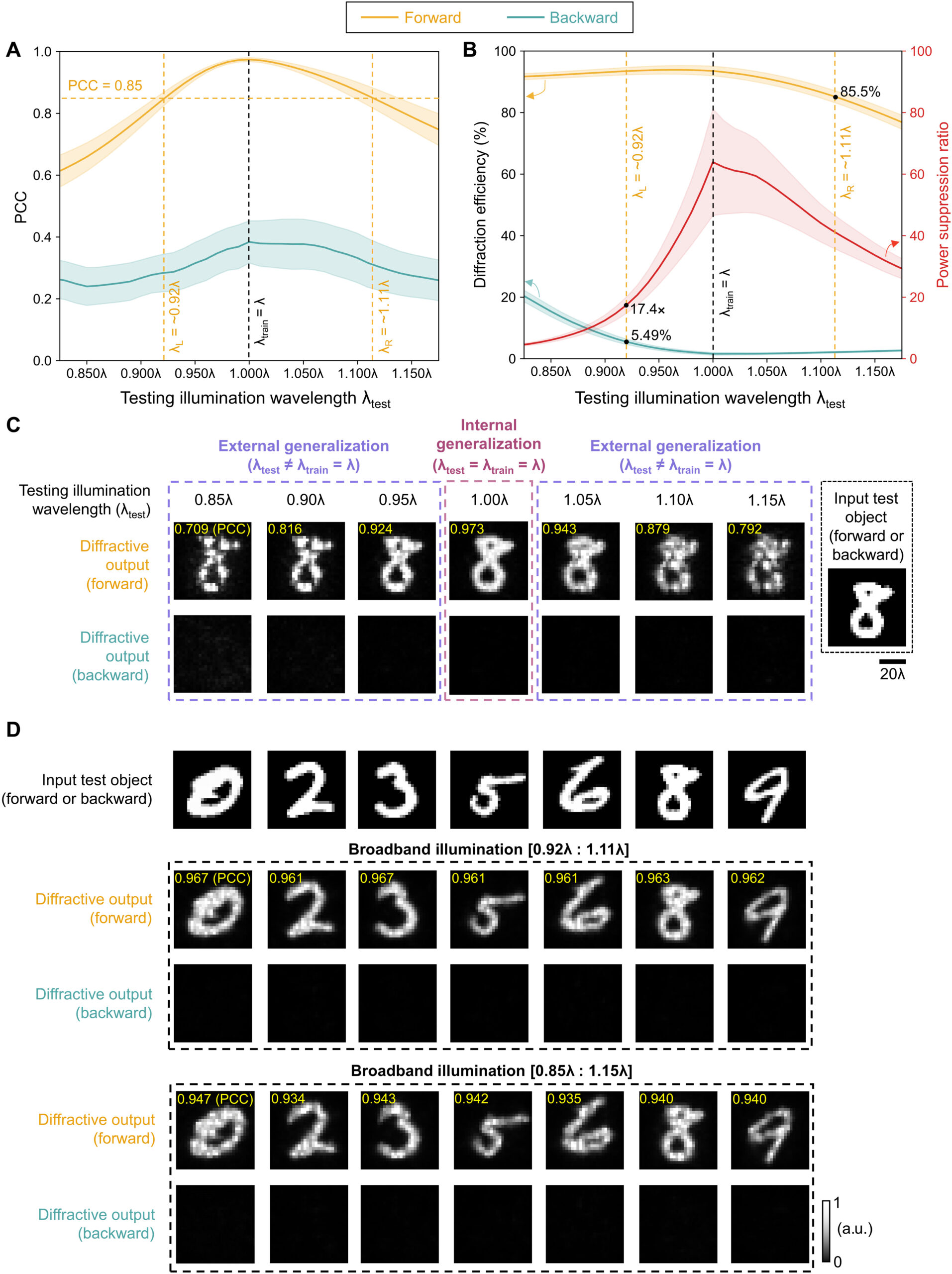
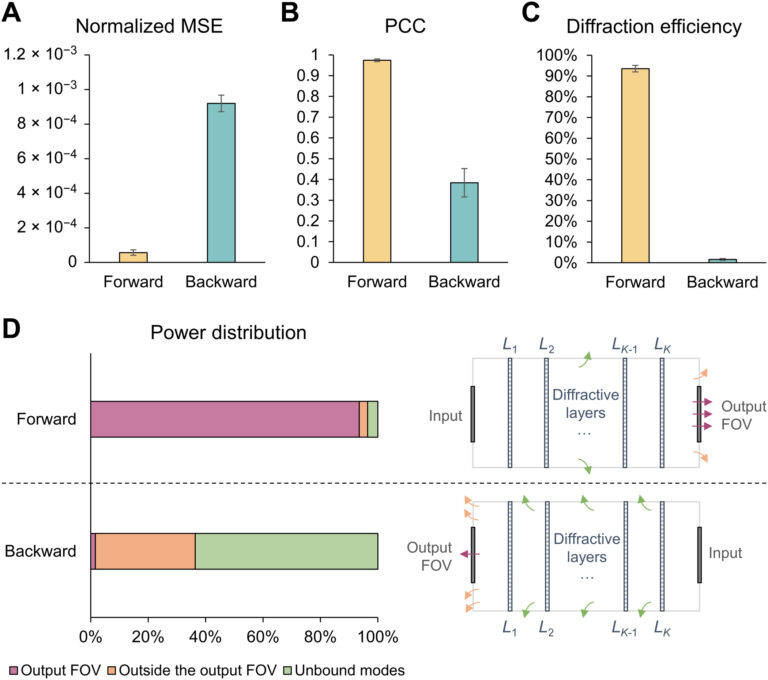
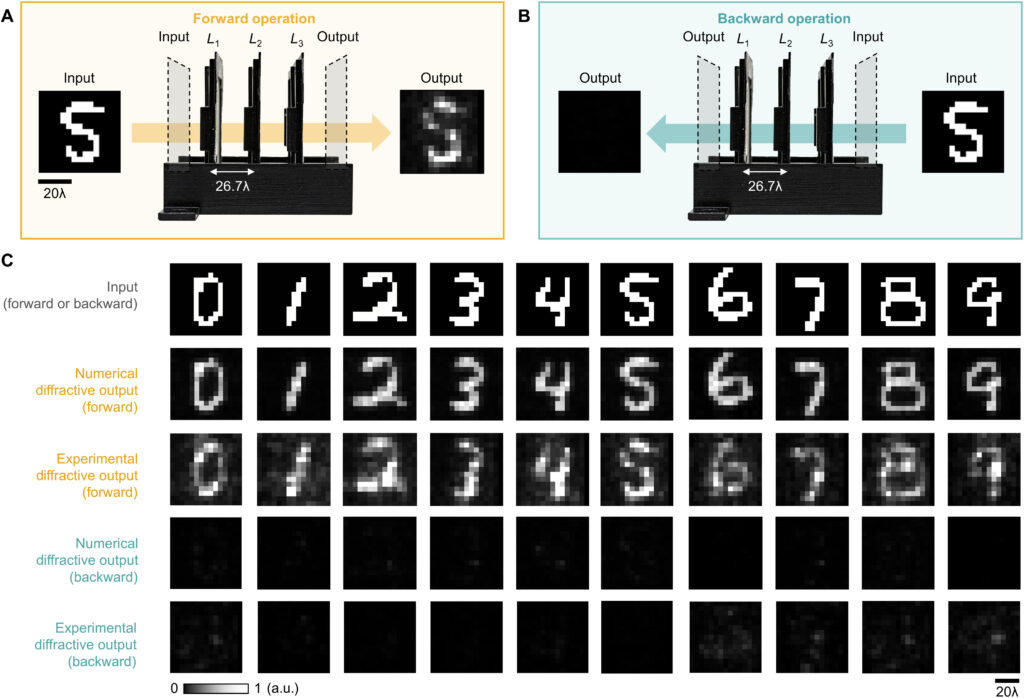
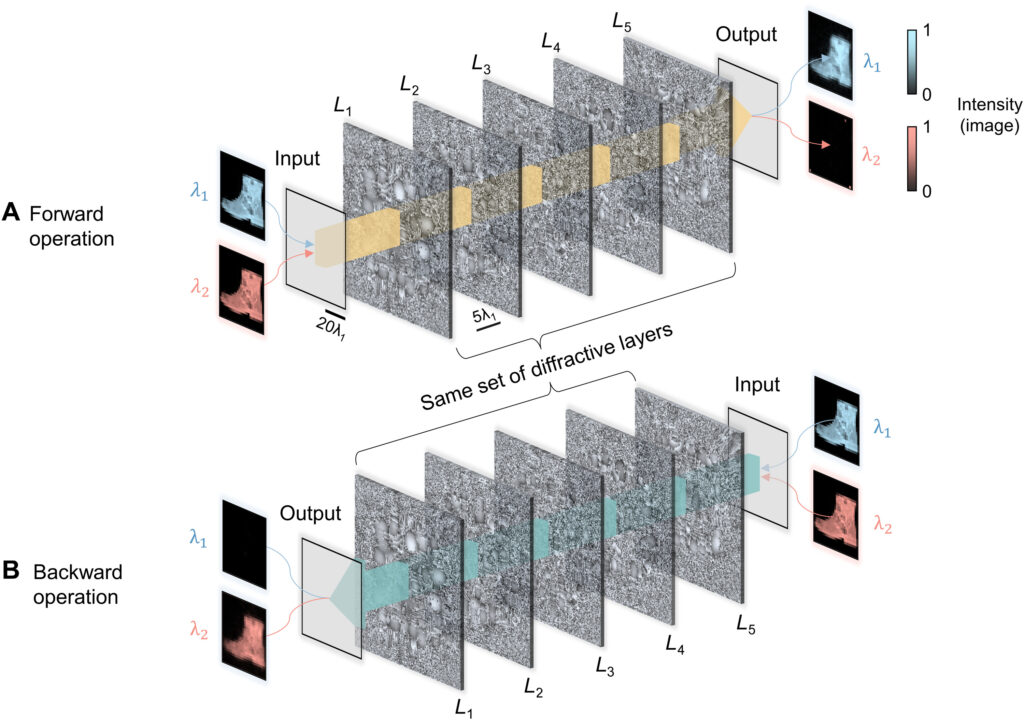
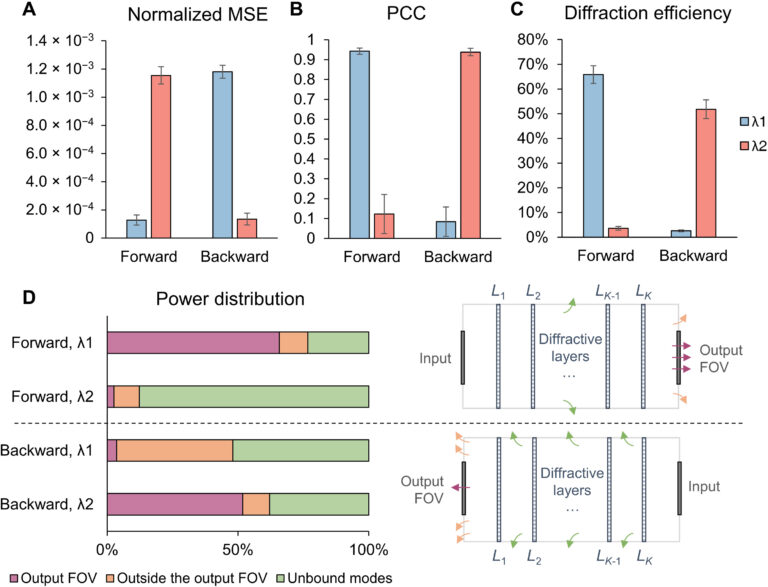
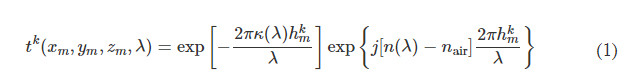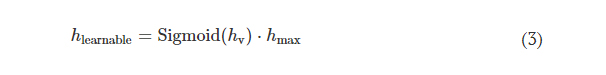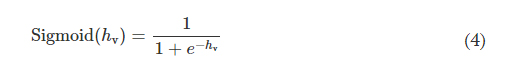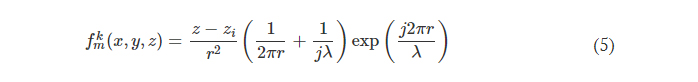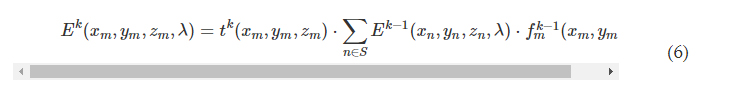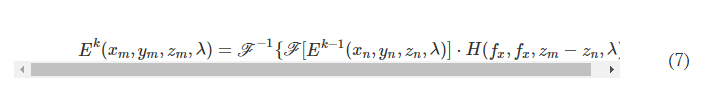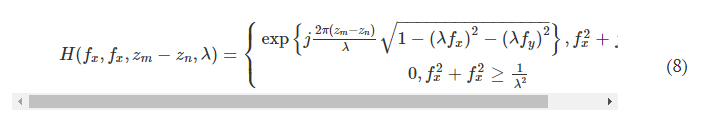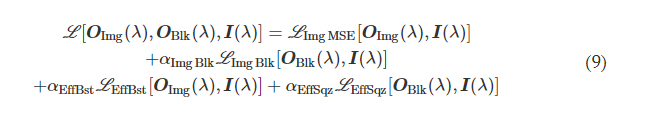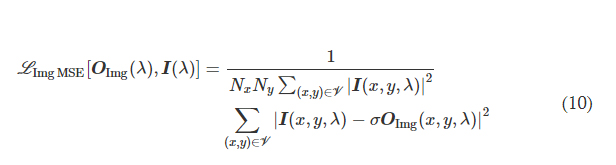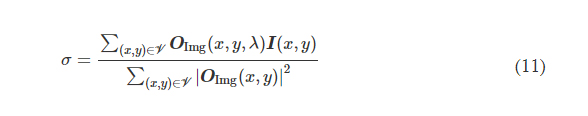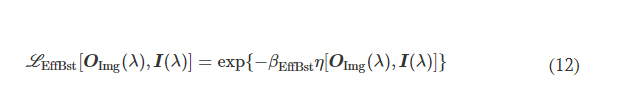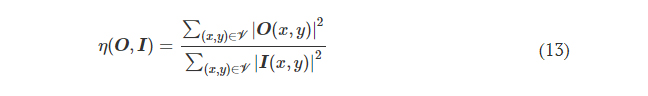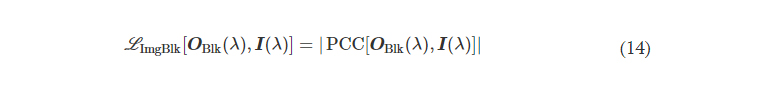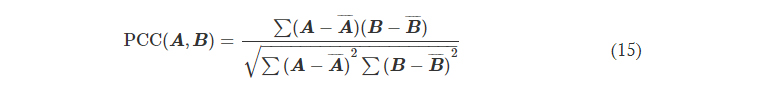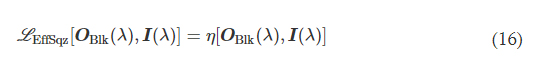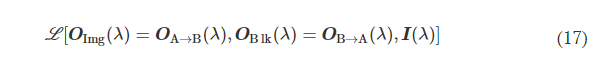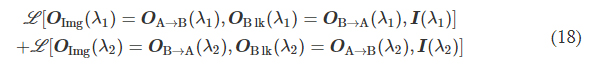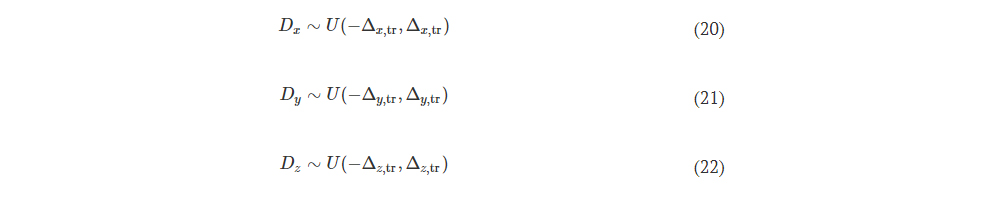Fig. 10. Performance analysis of the wavelength-multiplexed unidirectional diffractive imager shown in Fig. 9 and fig. S2. (A and B) Normalized MSE (A) and PCC (B) values calculated between the input images and their corresponding diffractive outputs at different wavelengths in the forward and backward operations. (C) The output diffraction efficiencies of the diffractive imager calculated in the forward and backward operations. In (A) to (C), the metrics are benchmarked across the entire MNIST test dataset and shown with their mean values and SDs added as error bars. (D) Left: The power of the different spatial modes at the two wavelengths propagating in the diffractive volume during the forward and backward operations, shown as percentages of the total input power. Right: Schematic of the different spatial modes propagating in the diffractive volume.
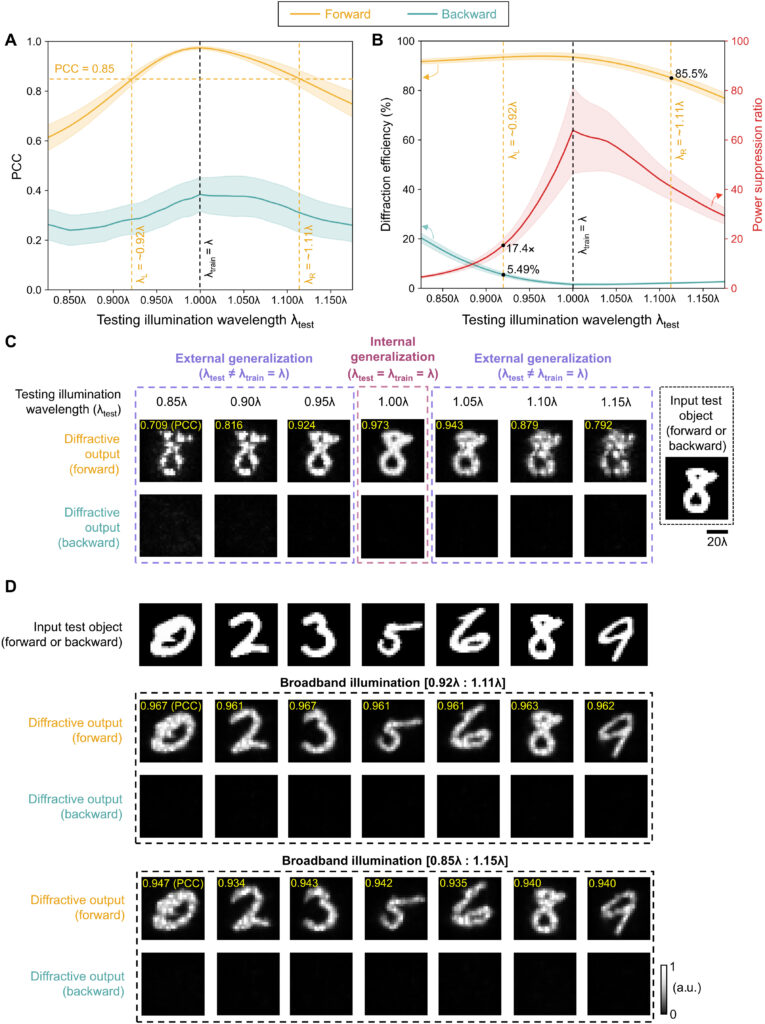
We should also note that, since this wavelength-multiplexed unidirectional imager was trained at two distinct wavelengths that control the opposite directions of imaging, the spectral response of the resulting diffractive imager, after its optimization, is vastly different from the broadband response of the earlier designs, reported in, e.g., Fig. 5. Figure S5 reveals that the wavelength-multiplexed unidirectional imager (as desired and expected) switches its spectral behavior in the range between λ1 and λ2, since its training aimed unidirectional imaging at opposite directions at these two predetermined wavelengths. Therefore, this spectral response that is summarized in fig. S5 is in line with the training goals of this wavelength-multiplexed unidirectional imager. However, it still maintains its unidirectional imaging capability over a range of wavelengths in both directions. For example, fig. S5 reveals that the output image PCC values for A → B remain ≥0.85 within the entire spectral range covered by 0.975 × λ1 to 1.022 × λ1 without any considerable increase in the diffraction efficiency for the reverse path, B → A. Similarly, the output image PCC values for B → A remain ≥0.85 within the entire spectral range covered by 0.968 × λ2 to 1.029 × λ2 without any noticeable increase in the diffraction efficiency for the reverse path, A → B, within the same spectral band. These results highlighted in fig. S5 indicate that the wavelength-multiplexed unidirectional imager can also operate over a continuum of wavelengths around λ1 (A → B) and λ2 (B → A), although the width of these bands are narrower compared to the broadband imaging results reported in Fig. 5.
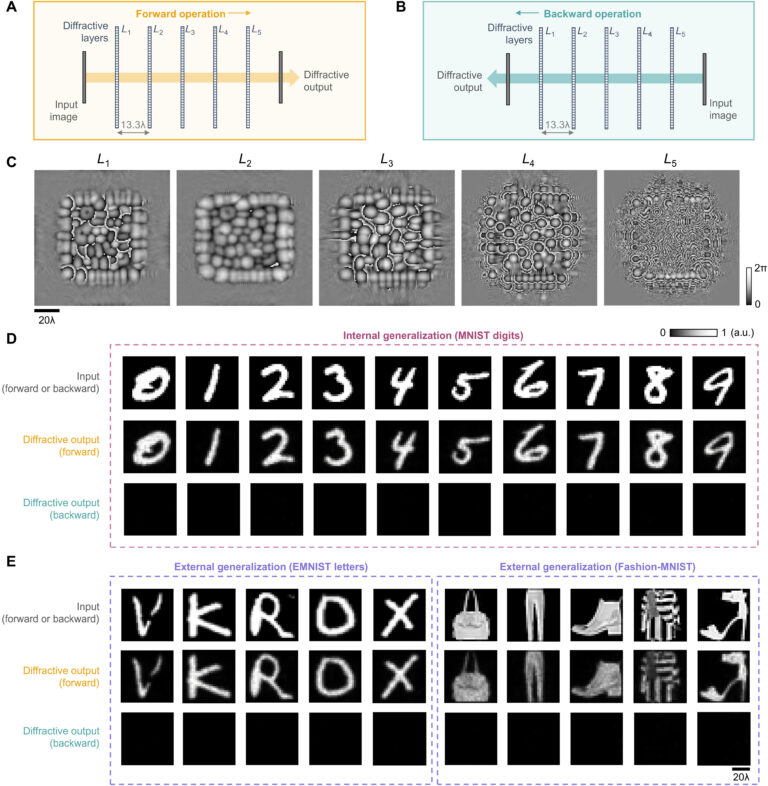
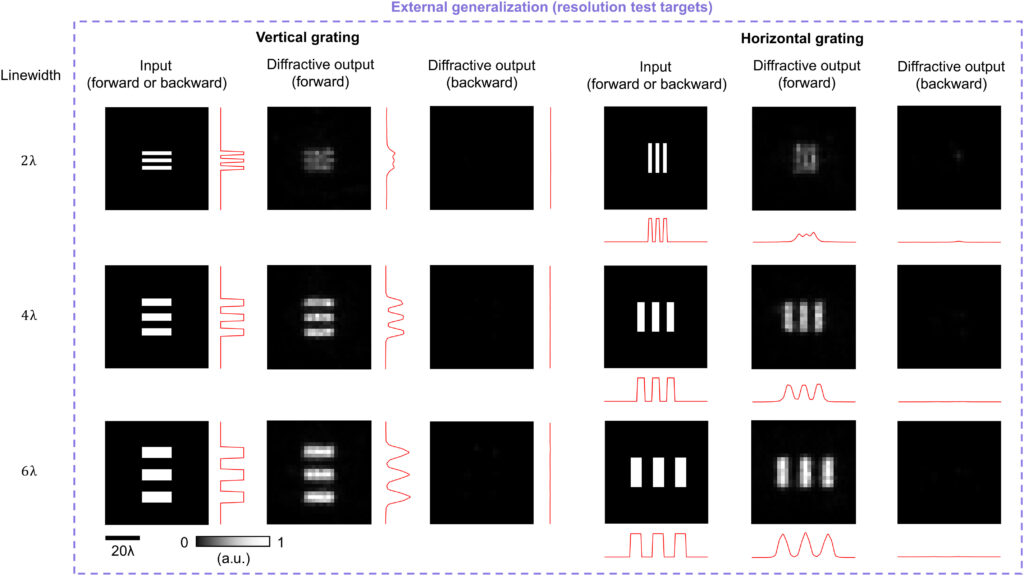
Last, we also tested the external generalization capability of this wavelength-multiplexed unidirectional imager on different datasets: handwritten letter images and fashion products as well as the contrast-reversed versions of these datasets. The corresponding imaging results are shown in Fig. 9D and fig. S6, once again confirming that our diffractive model successfully converged to a data-independent, generic imager where unidirectional imaging of various input objects can be achieved along either the forward or backward directions that can be switched/controlled by the illumination wavelength.
DISCUSSION
Our results constitute the first demonstration of unidirectional imaging. This framework uses structured materials formed by phase-only diffractive layers optimized through deep learning and does not rely on nonreciprocal components, nonlinear materials, or an external magnetic field bias. Because of the use of isotropic diffractive materials, the operation of our unidirectional imager is insensitive to the polarization of the input light, also preserving the input polarization state at the output. As we reported earlier in Results (Fig. 5), the presented diffractive unidirectional imagers maintain unidirectional imaging functionality under broadband illumination, over a large spectral band that covers, e.g., 0.85 × λ to 1.15 × λ, despite the fact that they were only trained using monochromatic illumination at λ. This broadband imaging performance was further enhanced, covering even larger input bandwidths, by training the diffractive layers of the unidirectional imager using a set of illumination wavelengths randomly sampled from the desired spectral band of operation as illustrated in fig. S7.
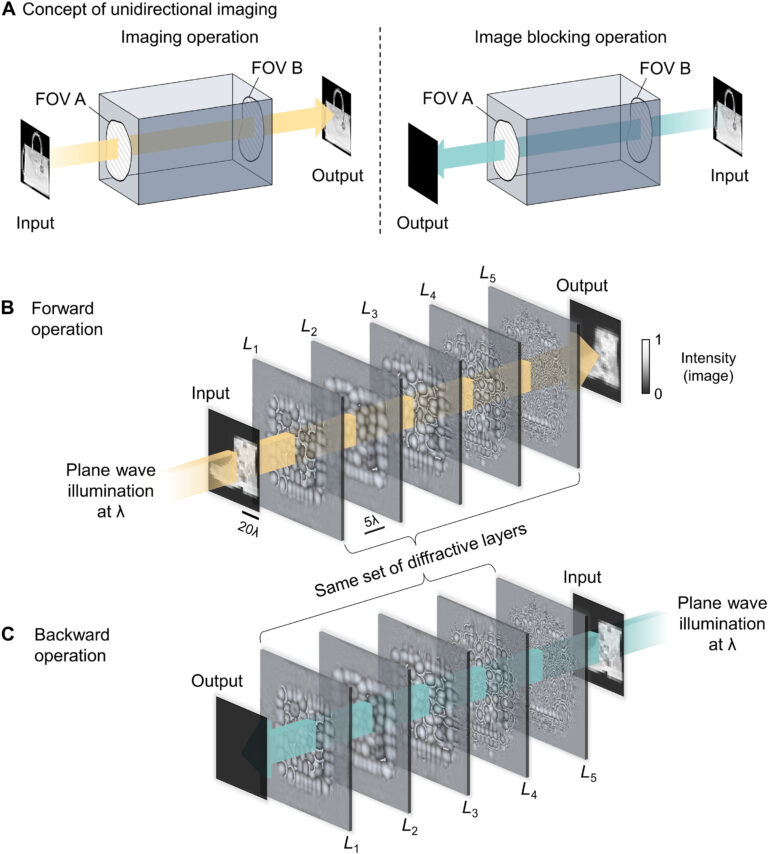
By examining the diffractive unidirectional imager design and the analyses shown in Fig. 2 and fig. S1, one can gain more insights into its operation principles from the perspective of the spatial distribution of the propagating optical fields within the diffractive imager volume. The diffractive layers L1 to L3 shown in Fig. 2C exhibit densely packed phase islands, similar to microlens arrays that communicate between successive layers. Conversely, the diffractive layers L4 and L5 have rapid phase modulation patterns, resulting in high spatial frequency modulation and scattering of light. Consequently, the propagation of light through these diffractive layers in different sequences leads to the modulation of light in an asymmetric manner (A → B versus B → A). To gain more insights into this, we calculated the spatial distributions of the optical fields within the diffractive imager volume in fig. S1 (C and D) for a sample object. We observe that, in the forward direction (A → B), the diffractive layers arranged with the order of L1 to L5 ensured that these optical fields propagated forward through the focusing by the microlens-like phase islands located in the diffractive layers L1 to L3, and as a result, the majority of the input power was maintained within the diffractive volume, creating a power efficient image of the input object at the output FOV. However, for the backward operation (B → A) where the diffractive layers are arranged in the reversed order (L5 to L1), the optical fields in the diffractive volume are initially modulated by the high spatial frequency phase patterns of the diffractive layers (i.e., L5 and L4), and during the early stages of the propagation within the diffractive volume, this leads to a large amount of radiation being channeled to the outer space aside the diffractive volume, in the form of unbound modes (see the green shaded areas in fig. S1, A and B). For the remaining spatial modes that managed to stay within the diffractive volume (propagating from B to A), they were guided by the subsequent diffractive layers (i.e., L3 to L1) to remain outside the output FOV (i.e., ending up within the orange shaded areas in fig. S1B).
One should note that the intensity distributions formed by these modes that lie outside the output FOV can be potentially measured by using, for example, side cameras that capture some of these scrambled modes. Such side cameras, however, cannot directly lead to meaningful, interpretable images of the input objects, as also illustrated in fig. S1. With the precise knowledge of the diffractive layers and their phase profiles and positions, one could potentially train a reconstruction digital neural network to make use of such side-scattered fields to recover the images of the input objects in the reverse direction of the unidirectional imaging system. This “attack” to digitally recover the lost image of the input object through side cameras and learning-based digital image reconstruction methods would not only require precise knowledge of the fabricated diffractive imager but can also be mitigated by surrounding the diffractive layers and the regions that lie outside the image FOV (orange regions in fig. 1, A and B) with absorbing layers/coatings that would protect the unidirectional imager against “hackers,” blocking the measurement of the scattered fields, except the output image aperture. Such absorbing layers also break the time-reversal symmetry of the imaging system, which help mitigate the risk of deciphering and decoding the original input in the backward direction.
Throughout this manuscript, we presented diffractive unidirectional imagers with input and output FOVs that have 28 by 28 pixels, and these designs were based on transmissive diffractive layers, each containing ≤200 by 200 trainable phase-only features. To further enhance the unidirectional imaging performance of these diffractive designs, one strategy would be to create deeper architectures with more diffractive layers, also increasing the total number (N) of trainable features. In general, deeper diffractive architectures present advantages in terms of their learning speed, output power efficiency, transformation accuracy, and spectral multiplexing capability (39, 44, 47, 48). Suppose an increase in the space-bandwidth product (SBP) of the input FOV A (SBPA) and the output FOV B (SBPB) of the unidirectional imager is desired, for example, due to a larger input FOV and/or an improved resolution demand; in that case, this will necessitate an increase in N proportional to SBPA × SBPB, demanding larger degrees of freedom in the diffractive unidirectional imager to maintain the asymmetric optical mode processing over a larger number of input and output pixels. Similarly, the inclusion of additional diffractive layers and features to be jointly optimized would also be beneficial for processing more complex input spectra through diffractive unidirectional imagers. In addition to the wavelength-multiplexed unidirectional imager reported in Figs. 8 to 10, an enhanced spectral processing capability through a deeper diffractive architecture may permit unidirectional imaging with, e.g., a continuum of wavelengths or a set of discrete wavelength across a desired spectral band. Furthermore, by properly adjusting the diffractive layers and the learnable phase features on each layer, our designs can be adapted to input and output FOVs that have different numbers and/or sizes of pixels, enabling the design of unidirectional imagers with a desired magnification or demagnification factor.
Although the presented diffractive unidirectional imagers are based on spatially coherent illumination, they can also be extended to spatially incoherent input fields by following the same design principles and deep learning–based optimization methods presented in this work. Spatially incoherent input radiation can be processed using phase-only diffractive layers optimized through the same loss functions that we used to design unidirectional imagers reported in our Results. For example, each point of the wavefront of an incoherent field can be decomposed, point by point, into a spherical secondary wave, which coherently propagates through the diffractive phase-only layers; the output intensity pattern will be the superposition of the individual intensity patterns generated by all the secondary waves originating from the input plane, forming the incoherent output image. However, the simulation of the propagation of each incoherent field through the diffractive layers requires a considerably increased number of wave propagation steps compared to the spatially coherent input fields, and as a result, the training of spatially incoherent diffractive imagers would take longer.
MATERIALS AND METHODS
Numerical forward model of a diffractive unidirectional imager
In the forward model of our diffractive unidirectional imager design, the input plane, diffractive layers, and output plane are positioned sequentially along the optical axis, where the axial spacing between any two of these layers (including the input and output planes) is set as d. For the numerical and the experimental models used here, the value of d is empirically chosen as 10 and 20 mm, respectively, corresponding to 13.33λ and 26.67λ, where λ = 0.75 mm. In our numerical simulations, the diffractive layers are assumed to be thin optical modulation elements, where the mth neuron on the kth layer at a spatial location (xm, ym, zm) represents a wavelength-dependent complex-valued transmission coefficient, tk, given by
where n(λ) and κ(λ) are the refractive index and the extinction coefficient of the diffractive layer material, respectively; these correspond to the real and imaginary parts of the complex-valued refractive index n~(λ) , i.e., n~(λ)=n(λ)+jκ(λ) (34). For the diffractive unidirectional imager validated experimentally at λ = 0.75 mm, the values of n~(λ) are measured using a terahertz spectroscopy system to reveal n(λ) = 1.700 and κ(λ) = 0.017 for the 3D printing material that we used. The same refractive index value n(λ) = 1.700 is also used in all the diffractive imager models used in our numerical analyses with κ = 0. hkm denotes the thickness value of each diffractive feature on a layer, which can be written as
where hlearnable refers to the learnable thickness value of each diffractive feature and is confined between 0 and hmax. The additional base thickness, hbase, is a constant that serves as the substrate (mechanical) support for the diffractive layers. To constrain the range of hlearnable, an associated latent trainable variable hv was defined using the following analytical form
where Sigmoid(hv) is defined as
Note that before the training starts, hv values of all the diffractive features were initialized as 0. In our implementation, hmax is chosen as 1.07 mm for the diffractive models that use λ = 0.75 mm so that the phase modulation of the diffractive features covers 0 to 2π. For the diffractive imager model that performs wavelength-multiplexed unidirectional imaging, hmax was empirically selected as 1.6 mm, still covering 0 to 2π phase range for both wavelengths (λ1 = 0.75 mm and λ2 = 0.85 mm). The substrate thickness, hbase, was assumed to be 0 in the numerical diffractive models and was chosen as 0.5 mm in the diffractive model used for the experimental validation. The diffractive layers of a unidirectional imager are connected to each other by free space propagation, which is modeled through the Rayleigh-Sommerfeld diffraction equation (33, 49)
where fkm(x,y,z,λ) is the complex-valued field on the mth pixel of the kth layer at (x, y, z), which can be viewed as a secondary wave generated from the source at (xm, ym, zm), r=(x−xm)2+(y−ym)2+(z−zm)2−−−−−−−−−−−−−−−−−−−−−−−−−−−√ , and j=−1−−−√ . For the kth layer (k ≥ 1, assuming that the input plane is the 0th layer), the modulated optical field Ek at location (xm, ym, zm) is given by
where S denotes all the diffractive features located on the previous diffractive layer. In our implementation, we used the angular spectrum approach (33) to compute Eq. 6, which can be written as
where F and F −1 denote the 2D Fourier transform and the inverse Fourier transform operations, respectively, both implemented using a fast Fourier transform. H(xn, yn, zm − zn, λ) is the transfer function of free space
where fx and fy represent the spatial frequencies along the x and y directions, respectively.
Training loss functions and image quantification metrics
We first consider a generic form of a diffractive unidirectional imager, where the image formation is permitted in one direction (e.g., A → B), and it is inhibited in the opposite direction (e.g., B → A) at a single training wavelength, λ. The training loss function for such a diffractive unidirectional imager was defined as
where I(λ) stands for the input image illuminated at a wavelength of λ and OImg(λ) and OBlk(λ) denote the output images in the forward and backward directions, respectively. All the input and output images have the perspective of the illumination beam direction, flipping them left to right as one switches the illumination direction, A → B or B → A. L ImgMSE penalizes the normalized MSE between the OImg(λ) and its ground truth, which can be written as
where *(x, y, λ) indexes the individual pixels at spatial coordinates (x, y) and wavelength λ and V denotes the defined FOV that has Nx × Ny pixels at the input or output plane. σ is a normalization constant used to normalize the energy of the diffractive output, thereby ensuring that the computed MSE value is not influenced by the errors arising from the output diffraction efficiency (50), and it is given by the following expression
L EffBst is used to improve the output diffraction efficiency along the imaging direction (e.g., A → B), which is defined as
where η(·) is the output diffraction efficiency of the diffractive unidirectional imager and βEffBst is an empirical weight coefficient, which was set as 1.0 during the training of all the diffractive models. η was defined as
L ImgBlk is defined to penalize the structural resemblance between the input image and the diffractive imager output along the image blocking direction (e.g., B → A)
where PCC stands for the Pearson correlation coefficient, defined as
L EffSqz in Eq. 9 is used to penalize the output diffraction efficiency in the backward direction
αImgBlk, αEffBst, and αEffSqz in Eq. 9 are the empirical weight coefficients associated with L ImgBlk, L EffBst, and L EffSqz, respectively. We denote the diffractive unidirectional imager output images for A → B and B → A as OA→B(λ) and OB→A(λ), respectively. For the diffractive unidirectional imaging models that were trained using a single illumination wavelength (e.g., in Figs. 2 and 7), the image formation is set to be maintained in the forward direction (A → B) and inhibited in the backward direction (B → A), i.e., OImg(λ) = OA→B(λ) and OBlk(λ) = OB→A(λ). Therefore, the loss function for training these models can be formulated as
where L (·) refers to the same loss function defined in Eq. 9. During the training of the unidirectional imager models with five diffractive layers and a single training wavelength channel, the empirical weight coefficients αImgBlk, αEffBst, and αEffSqz were set as 0, 0.001, and 0.001, respectively; during the training of the other model with three diffractive layers used for the experimental validation, the same weight coefficients were set as 0, 0.01, and 0.003, respectively.
For the wavelength-multiplexed unidirectional diffractive imager model shown in Fig. 9, at λ1, the image formation is permitted in the direction A → B and inhibited in the direction B → A, whereas at λ2, the image formation is permitted in the direction B → A and inhibited in the direction A → B, respectively; i.e., OImg(λ1) = OA→B(λ1), OBlk(λ1) = OB→A(λ1), OImg(λ2) = OB→A(λ2), and OBlk(λ2) = OA→B(λ2). Accordingly, we formulated the loss function used for training this model as
where L (·) refers to the loss function defined in Eq. 9. During the training of this model, the weight coefficients αImgBlk, αEffBst, and αEffSqz were empirically set as 0.0001, 0.001, and 0.001, respectively.
For quantifying the imaging performance of the presented diffractive imager designs, the reported values of the output MSE, output PCC, and output diffraction efficiency were directly taken from the calculated results of L ImgMSE, PCC, and η, respectively, revealing the averaged values across the blind testing image dataset. When calculating the power distributions of different optical modes within the diffractive volume, the power percentage of the output FOV modes takes the same value as η, and the power percentage outside the output FOV is computed by subtracting the total power integrated within the output image FOV from the total power integrated across the entire output plane. The power in the absorbed modes is calculated by summing up the power loss before and after the optical field modulation by each diffractive layer. After excluding the power of the above modes from the total input power, the remaining part is calculated as the power of the unbound modes.
Training details of the diffractive unidirectional imagers
For the numerical models used here, the smallest sampling period for simulating the complex optical fields is set to be identical to the lateral size of the diffractive features, i.e., ~0.53λ for λ = 0.75 mm. The input/output FOVs of these models (i.e., FOV A and B) share the same size of 44.8 by 44.8 mm2 (i.e., ~59.7λ × 59.7λ) and are discretized into 28 by 28 pixels, where an individual pixel corresponds to a size of 1.6 mm (i.e., ~2.13λ), indicating a four-by-four binning performed on the simulated optical fields.
For the diffractive model used for the experimental validation of unidirectional imaging, the sampling period of the optical fields and the lateral size of the diffractive features are chosen as 0.24 and 0.48 mm, respectively (i.e., 0.32λ and 0.64λ). This also results in a two-by-two binning in the sampling space where an individual feature on the diffractive layers corresponds to four sampling space pixels that share the same dielectric material thickness value. The input and output FOVs of this model (i.e., FOV A and B) share the same size of 36 by 36 mm2 (i.e., 48λ × 48λ) and are sampled into arrays of 15 by 15 pixels, where an individual pixel has a size of 2.4 mm (i.e., 3.2λ), indicating that a 10-by-10 binning is performed at the input/output fields in the numerical simulation.
During the training process of our diffractive models, an image augmentation strategy was also adopted to enhance their generalization capabilities. We implemented random translation, random up-to-down, and random left-to-right flipping of the input images using the transforms.RandomAffine function built-in PyTorch. The translation amount was uniformly sampled within a range of [−10, 10] and [−5, 5] pixels in the diffractive unidirectional imager models used for numerical analysis and the model used for the experimental validation, respectively. The flipping operation is set to be performed at a probability of 0.5.
All the diffractive imager models used in this work were trained using PyTorch (v1.11.0, Meta Platforms Inc.). We selected AdamW optimizer (51, 52), and its parameters were taken as the default values and kept identical in each model. The batch size was set as 32. The learning rate, starting from an initial value of 0.03, was set to decay at a rate of 0.5 every 10 epochs, respectively. The training of the diffractive models was performed with 50 epochs. For the training of our diffractive models, we used a workstation with a GeForce GTX 1080Ti graphical processing unit (Nvidia Inc.) and Core i7-8700 central processing unit (Intel Inc.) and 64 GB of RAM, running Windows 10 operating system (Microsoft Inc.). The typical time required for training a diffractive unidirectional imager is ~3 hours.
Vaccination of the diffractive unidirectional imager against experimental misalignments
During the training of the diffractive unidirectional imager design for experimental validation, possible inaccuracies imposed by the fabrication and/or mechanical assembly processes were taken into account in our numerical model by treating them as random 3D displacements (D) applied to the diffractive layers (53). D can be written as
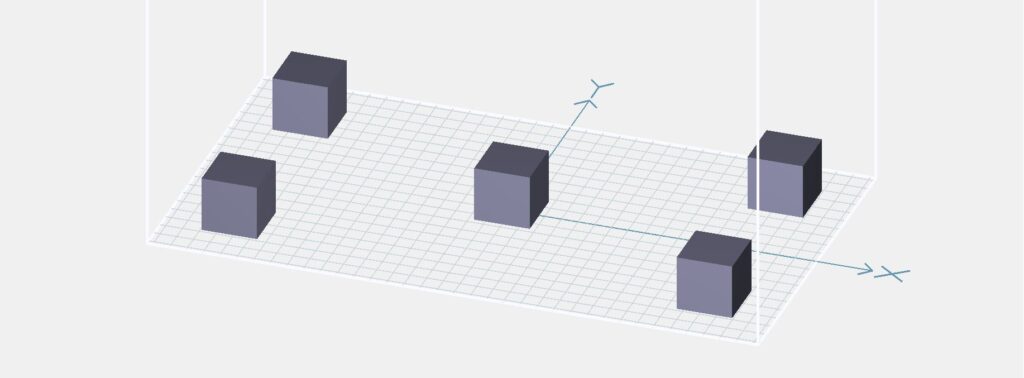
where Dx and Dy represent the random lateral displacement of a diffractive layer along the x and y directions, respectively, and Dz represents the random perturbation added to the axial spacing between any two adjacent layers (including diffractive layers, input FOV A, and output FOV B). Dx, Dy, and Dz of each diffractive layer were independently sampled based on the following uniform (U) random distributions
where Δ*,tr denotes the maximum amount of shift allowed along the corresponding axis, which was set as Δx,tr = Δy,tr = 0.48 mm (i.e., 0.64λ) and Δz,tr = 1.5 mm (i.e., 2λ) during the training process. Following the training under this vaccination strategy, the resulting diffractive unidirectional imager shows resilience against possible misalignments in the fabrication and assembly of the diffractive layers.
Note that, in addition to the 3D displacements of the diffractive layers, there may also exist other types of alignment errors in our experimental setup, such as 3D rotational misalignments of the diffractive layers. However, since the holders used to fix the diffractive layers are, in general, manufactured with high structural precision and surface flatness, we did not incorporate these types of misalignments into our forward model considering their negligible impact in our case. In the event that such rotational misalignments of the diffractive layer become an important factor in the experimental results, the undesired in-plane rotations of the diffractive layers can be readily modeled through applying a 2D coordinate transformation based on unitary rotation matrices, while the out-of-plane rotation of the diffractive layers can be addressed by modifications to the formulation of the wave propagation between tilted diffractive planes (53–55).
Experimental terahertz imaging setup
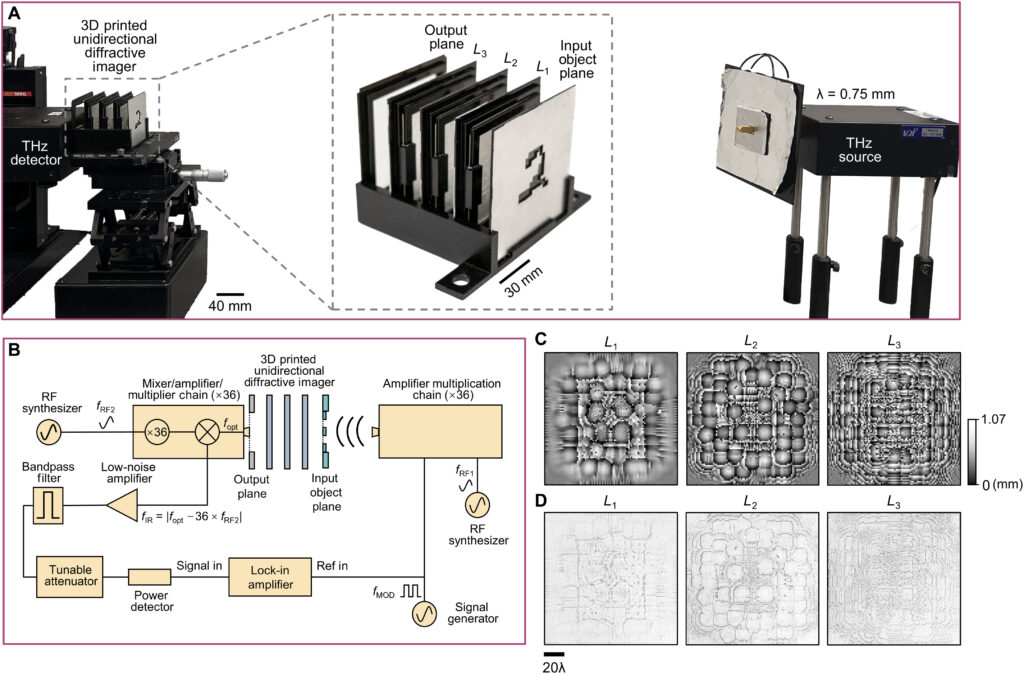
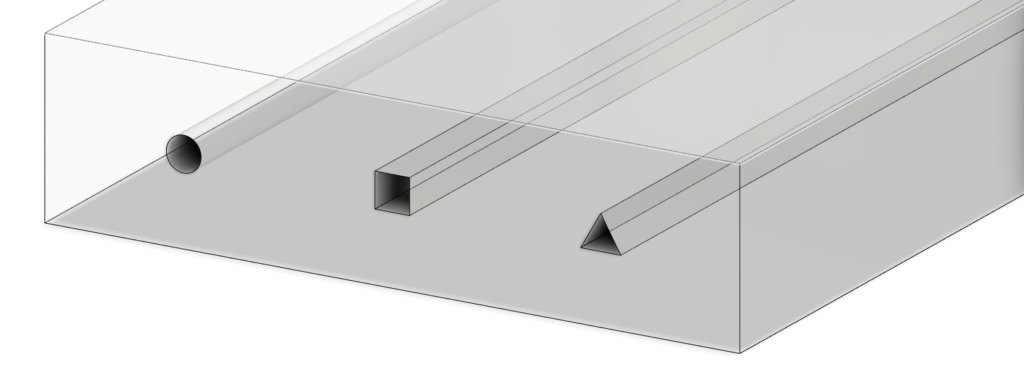
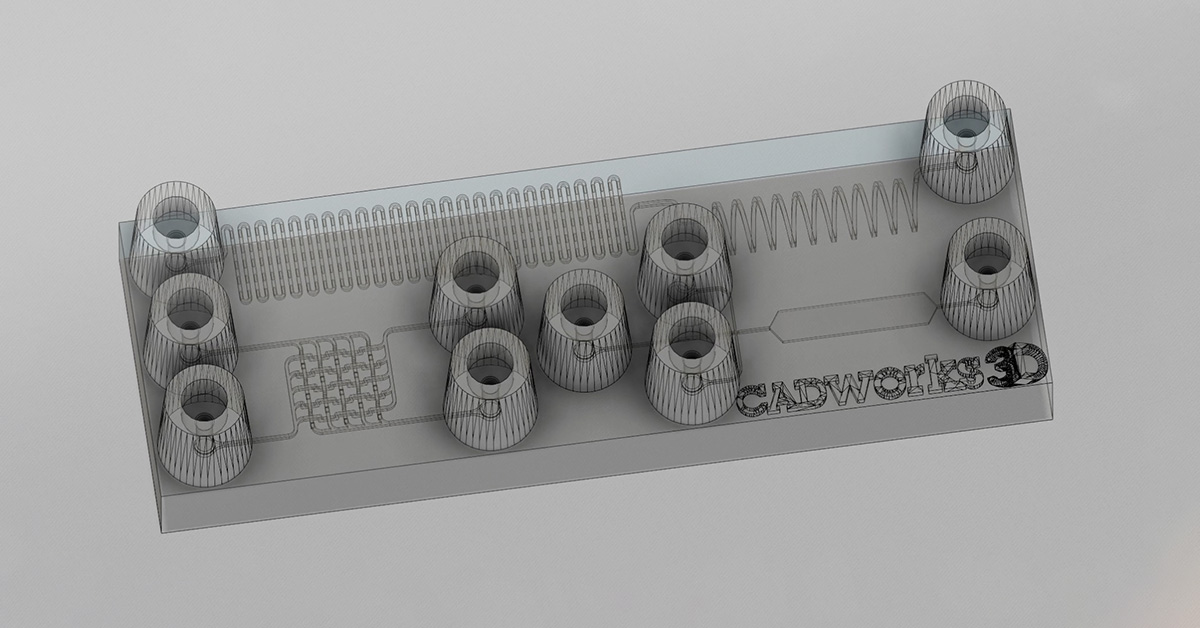
We fabricated the diffractive layers using a 3D printer (PR110, CADworks3D). The test objects were also 3D printed (Objet30 Pro, Stratasys) and coated with aluminum foil to define the light-blocking areas, with the remaining openings defining the transmission areas. We used a holder that was also 3D printed (Objet30 Pro, Stratasys) to assemble the printed diffractive layers along with input objects, following the relative positions of these components in our numerical design.
A terahertz continuous-wave scanning system was used for testing our diffractive unidirectional imager design. According to the experimental setup illustrated in Fig. 6B, we used a terahertz source in form of a WR2.2 modular amplifier/multiplier chain (AMC), followed by a compatible diagonal horn antenna (Virginia Diodes Inc.). A 10-dBm radiofrequency (RF) input signal at 11.1111 GHz (fRF1) at the input of AMC is multiplied 36 times to generate the output radiation at 400 GHz, corresponding to a wavelength of λ = 0.75 mm. The AMC output was also modulated with a 1-kHz square wave for lock-in detection. The assembled diffractive unidirectional imager is placed ∼600 mm away from the exit aperture of the horn antenna, which results in an approximately uniform plane wave impinging on its input FOV (A) with a size of 36 by 36 mm2 (i.e., 48λ × 48λ). The intensity distribution within the output FOV (B) of the diffractive unidirectional imager was scanned at a step size of 1 mm by a single-pixel mixer/AMC (Virginia Diodes Inc.) detector on an xy positioning stage that was built by combining two linear motorized stages (Thorlabs NRT100). The detector also receives a 10-dBm sinusoidal signal at 11.083 GHz (fRF2) as a local oscillator for mixing to down-convert the output signal to 1 GHz. The signal is then fed into a low-noise amplifier (Mini-Circuits ZRL-1150-LN+) with a gain of 80 dBm, followed by a band-pass filter at 1 GHz (± 10 MHz) (KL Electronics 3C40-1000/T10-O/O), so that the noise components coming from unwanted frequency bands can be mitigated. Then, after passing through a tunable attenuator (HP 8495B) used for linear calibration, the final signal is sent to a low-noise power detector (Mini-Circuits ZX47-60). The detector output voltage is measured by a lock-in amplifier (Stanford Research SR830) with the 1-kHz square wave used as the reference signal. Last, the lock-in amplifier readings were calibrated into a linear scale. In our postprocessing, linear interpolation was applied to each measurement of the intensity field to match the pixel size of the output FOV (B) used in the design phase, resulting in the output measurement images shown in Fig. 7C.












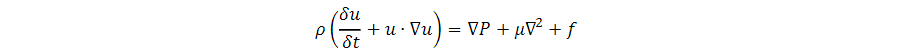




















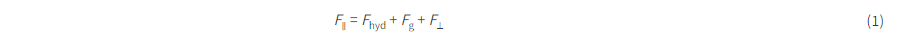












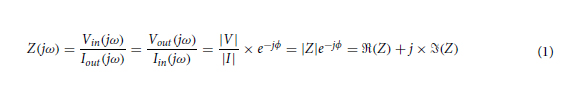
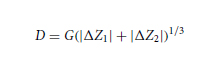






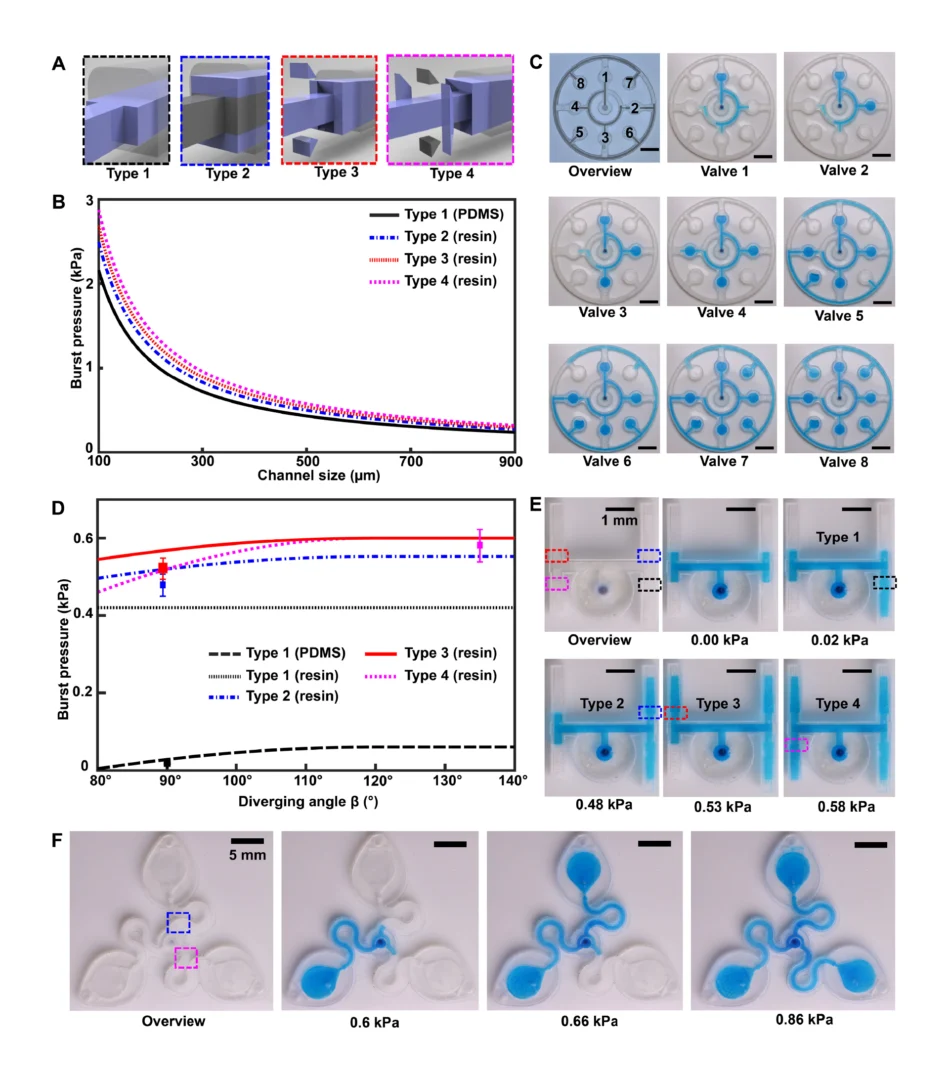
![Figure 2. Optimized design strategy for fabricating 3D printed epifluidic devices with prescribed channel geometries. (A) Photograph of 3D printed test channels [100 to 900 μm, square; 2-s layer cure time (LCT)]. (B) Plot of variation of printed channel height from designed dimensions as a function of LCT. (C) Plot of variation of printed channel width from designed dimensions as a function of LCT. (D) Plot highlighting the printable region of the digital light processing (DLP) printer used in this work for various channel dimensions relevant to epifluidic devices.](https://cadworks3d.com/wp-content/uploads/2025/07/Academic-Article_UH-Manoa_Skin-interfaced-microfluidic-systems-with-spatially-engineered-3D-fluidics-for-sweat-capture-and-analysis_Figure-2-430x1080.webp)